 Air pollution in London is increasing, and since 2010 toxic air has been at illegal levels in the capital. The extensive use of diesel engines, together with the burning of other fossil fuels, has been linked to increase particular matter above the considered safety level. What affect does urban air have on fungal spores abundance and diversity, how does this compare to rural, less polluted locations outside of the London haze, and are there frequency differences of known antifungal resistance alleles? To help answer these questions, we have been monitoring the abundance of Aspergillus fumigatus spores (plus others) in the the air above St Mary's hospital using a Burkard spore trap. The Burkard spore trap is a self-contained vacuum pump designed to sample air-borne particles, such as fungal spores, continuously for periods up to seven days.  Every Wednesday a party of intrepid explorers head onto the roof of the Medical School building where the Burkard trap sits, symbolically facing the windows of Sir Alexander Fleming's famous lab. Once collected the canister containing a weeks worth of fungal spores (amongst other things...) is shipped off to Rothamsted Research, home to the oldest, continuous agronomic experiments in the world (plus loads of other amazing research) . Here, the DNA from the fungal spores that has landed onto the strip will be extracted. Once the DNA is purified, a string of downstream analyses will be used to help us answer questions related to fungal spore diversity and antifungal resistance frequency in the Big Smoke. Keep your eye out for future updates!
3 Comments
 Archived soil samples collected by the Centre of Ecology and Hydrology Archived soil samples collected by the Centre of Ecology and Hydrology Hello and apologies for not introducing myself sooner- I’m Jenny and in October I joined Thomas and Johanna in the Evolution of Resistance group as a PhD student. My PhD started off with a whistle stop tour of the world of medical mycology when I attended the 8th Trends in Medical Mycology conference in Belgrade, Serbia. There I met many of the leading names in medical mycology and heard of the fascinating research being done on important human fungal pathogens, such as Aspergillus fumigatus and Candida auris, and how knowledge is being integrated into clinical management of the illnesses they cause. I was able to put names to the faces of several people I hope to collaborate with during the course of my PhD and I returned from the conference full of enthusiasm and ideas for my research proposal! For my project I will be looking at the background levels of Aspergillus fumigatus across South England and how these differ by rural and urban environments, season, land use and fungicide spraying. This will involve collecting plenty of air and soil samples from difficult locations and at varying times throughout the year and directly culturing Aspergillus, as well as extracting DNA for qPCR and sequencing analysis. Today is exciting because I received my first soil samples, collected as part of an ongoing project based at the Centre of Ecology and Hydrology (CEH) in Wallingford, Oxfordshire. These will form the basis of my pilot field study to see if I can culture and extract DNA from Aspergillus fumigatus in surface soil that has been frozen for long periods of time. My supervisors are Prof. Matthew Fisher of the Department of Infectious Disease Epidemiology at Imperial College London (ICL), Dr Andrew Singer of CEH, and Dr Marta Blangiardo of the Department of Epidemiology and Biostatistics at ICL. As mentioned in a previous post, my project is funded by NERC and is part of the Science and Solutions for a Changing Planet (SSCP) DTP run by Imperial’s Grantham Institute (so many acronyms!). I am fortunate also to have Public Health England (PHE) as my project CASE partner and will have the opportunity to work in their labs in my second or third year. Prior to starting my PhD I spent 2.5 years working as a Research Assistant in Mat Fisher’s group researching fungal diversity on amphibian skin and how this might protect them from Batrachochytrium dendrobatidis- a disease-causing chytrid fungi that causes morbidity and mortality in amphibians and has devastated frog populations worldwide. I am hoping the many skills I learnt in this role relating to fieldwork, labwork and data analysis will stand me in good stead for my project- in particular knowing that things CAN and WILL go wrong, especially in lab work, so to maintain infinite patience and not lose faith! I look forward to sharing more as my project progresses.  MARDy is a curated database of known resistance mechanisms and associated antifungal drugs that is managed by Fisher Lab members. MARDy currently includes amino acid substitutions, tandem repeat gene sequences, and chromosome ploidy. MARDy is the brain-child of Dr Joanna Rhodes and was developed in collaboration with Dr Anthony Nash (University of Oxford). Our job here at Fisher Lab is to maintain the database, introducing new searchable (and BLASTable!) entries as they are discovered. This will aid researchers and clinicians to identify mechanisms of resistance in environmental or medically important fungal strains. MARDy has its own twitter account, that can be used to keep up to date with novel entries and developmental updates. Publication is under review and the website is available for use now! If you have any questions please contact us here on via the MARDy twitter account. Things are continuing to progress nicely here at Imperial! We have just finalised our next batch of 96 isolates, which will be shipped off to the sequencer in a matter of days. These isolates have an environmental origin and are resistant to azole antifungals. They have a vast global distribution and are temporally very interesting. The genome sequences will help fill in the gaps on our phylogeny and provide us with more information on the ecology and evolution of the pathogen and its resistance phenotype. In addition to this, we have received more isolates from collaborators across the globe all of which will be included in our later WGS runs in 2018.
In other news, Jennifer Shelton has just started her PhD with us and will be investigating whether Aspergillus fumigatus distribution in air and soil is linked to environmental factors and clinical disease. The project is out of Imperial College but she will split her time between here and the Centre for Hydrology and Ecology in Wallingford. The project is part of the Science and Solutions for a Changing Planet (SSCP) Doctoral Training Program (DTP), which is funded by the Natural Environment Research Council (NERC) and Imperial’s Grantham Institute. You will hear from Jen soon so check back for updates on her project as well as our WGS runs. 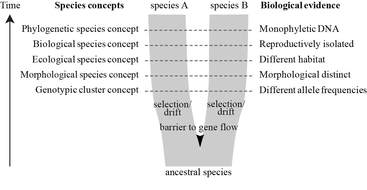 frederikleliaert.files.wordpress.com frederikleliaert.files.wordpress.com Over the years, many papers have attempted to prise open the secrets surrounding A. fumigatus population structure, with most papers falling upon an alternative conclusion. Probably the most notable, Pringle et al (2005), used five loci to identify genetic isolation between two well supported clades and concluded that A. fumigatus actually consists of two cryptic species, Aspergillus "fumigatus" fumigatus and Aspergillus "occultum" fumigatus. Pringle's observation were determined according to the phylogenetic species concept (PSC*), as opposed to the more commonly applied biological species concept or ecological species concept. At the same time, Rydholm et al (2005) investigated a collection of environmental A. fumigatus isolates at three intergenic loci for insights into population structure. In their paper, a comparison was made with closely related Neosartorya spp, N. fischeri and N. spinosa (Neosartorya spp. are the sexual teleomorphic stage of Aspergillus spp.). In accordance with a previous study, which used restriction fragment length polymorphism (Debeaupuis et al. 1997), Rydholm detected no evidence for population differentiation or sub-structure and concluded that lack of population structure could be mediated by large scale gene flow across continents, mostly due to the long distance dispersal of airborne A. fumigatus conidia. 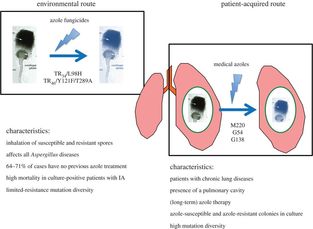 Meis et al 2016 Meis et al 2016 In recent years there has been renewed interest in determining population structure in A. fumigatus, motivated primarily by the emergence of global azole resistance in immunocompromised aspergillosis patients. Klassen et al (2012) and Ashu et al (2017), used microsatellites (a well established genetic marker often used in molecular ecology) to make conclusions on the genetic structure of both environmental and clinical A. fumigatus isolates, with and without azole resistance. Klassen investigated 255 isolates collected from the Netherlands, 25 of which had the azole resistant allele TR34/L98H (see previous post). Using two well established tools for determining genetic clustering (STRUCTURE** and DAPC), five genetically different, predominantly asexual populations were identified, with the azole resistance allele (TR34/L98H) found in just one of the five populations. Klassen concludes that confinement to a single asexual population suggests that sexual reproduction is not facilitating the emergence or spread of azole resistance, and that reproductive mode and genetic differentiation contribute to the structure of A. fumigatus populations in the Netherlands. Most recently, Ashu investigated a collection of 2026 globally distributed isolates. They found that their collection fit into eight genetic clusters, with limited but statistically significant geographic or ecological differentiation; somewhat confirming previous observations of ubiquity and long distant dispersion. Notably on this occasion, they found signatures of both asexual and sexual reproduction, with the TR34/L98H resistance allele found in more than just one population. Although fairly informative, microsatellite markers paint just part of the picture, encompassing a fraction of the genome. Our goal, as a genomics lab, is to unsurprisingly, analyse genomes. We are revisiting the story; firstly, by using information gleaned from the large collection of global A. fumigatus isolates genotyped by microsatellite markers, we have selected spatially and temporally important isolates for whole genome sequence analysis. We will then use the detailed information provided by genome sequencing and multiple sequence alignments to investigate genetic diversity, population structure, azole resistance evolution and mutation rates, to name but a few. Keep checking back for more updates and information regarding the labs progress and/or more information on the evolution of azole resistance in Aspergillus fumigatus.
* Speciation under the PSC can be defined as a group of organisms whose members are all more closely related to each other then they are to other organism outside of the groups. Membership is determined by nucleic acid variation and often visualised using multi-locus phylogenetic trees. It has been tremendously important in the identification of microbial species that are unculturable - especially fungi. N.B. blog post structure similar to our review - Untangling Aspergillus fumigatus population structure - Sewell et al (in writing) Further reading Pringle, A., Baker, D. M., Platt, J. L., Wares, J. P., Latge, J. P., & Taylor, J. W. (2005). Cryptic speciation in the cosmopolitan and clonal human pathogenic fungus Aspergillus fumigatus. Evolution, 59(9), 1886-1899. Rydholm, C., Szakacs, G. and Lutzoni, F., (2006). Low genetic variation and no detectable population structure in Aspergillus fumigatus compared to closely related Neosartorya species. Eukaryotic cell, 5(4), pp.650-657. Debeaupuis, J.P., Sarfati, J., Chazalet, V. and Latge, J.P., 1997. Genetic diversity among clinical and environmental isolates of Aspergillus fumigatus. Infection and Immunity, 65(8), pp.3080-3085. Klaassen, C.H., Gibbons, J.G., Fedorova, N.D., Meis, J.F. and Rokas, A., (2012). Evidence for genetic differentiation and variable recombination rates among Dutch populations of the opportunistic human pathogen Aspergillus fumigatus. Molecular ecology, 21(1), pp.57-70. Ashu, E. E., Hagen, F., Chowdhary, A., Meis, J. F., & Xu, J. (2017). Global Population Genetic Analysis of Aspergillus fumigatus. MSphere, 2(1), e00019-17.  Over the last few weeks we have been collecting more Aspergillus fumigatus isolates from around the world to add to our ever-expanding collection. These isolates are of interest both spatially and temporally and will be used to answer some fundamental questions related to the evolution of global antifungal drug resistance. Some of these strains have been sent to us by the good folks at Westerdijk Fungal Biodiversity Institute (check them out). We have got some fascinating specimens that will soon be heading off to Edinburgh for WGS. In addition to these interesting isolates, we now have a number of airborne specimens collected from Hyde Park and St Mary's Hospital roof. These, along with other samples collected around London, will be genotyped for resistance in due course and potentially added to a later WGS-ing run. Progress is moving along nicely and a neat set of clinically important isolates will soon be loaded onto the sequencer. Sequence data from our previous run is churning through our bioinformatics pipeline. And our new isolates are happily growing in our incubators. Please continue to check back here for updates on our progress and if you have any question, please feel free to get in touch! 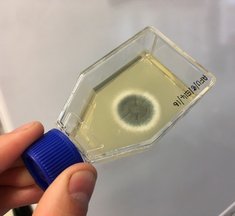 An Aspergillus fumigatus colony. An Aspergillus fumigatus colony. In the age of next generation sequencing, research on antimicrobial resistance evolution can be done practically using whole genome sequencing (WGS). From the identification of novel resistance mechanisms to spatially tracking the evolution and transmission of resistance alleles, the whole genome approach is fast becoming an invaluable tool in the fight against the antimicrobial resistance crisis. Our goal is to use WGS - by sequencing and analysing ~400 Aspergillus fumigatus genomes from across the globe - to help us research the evolution of antifungal resistance. Antifungal resistance in A. fumigatus, which is fast becoming a credible threat to human health worldwide, is thought to have evolved in both the clinical and environmental setting. This is due to the nature of A. fumigatus; an environmental saprophyte but also a serious fungal pathogen to immunocompromised hosts, where it causes a range of symptoms from mild allergic reactions to invasive aspergillosis of the lungs. Azole antifungals are one of a few classes of drugs available for the treatment and prophylaxis of aspergillosis. Predictably however, after extensive use, resistance to azoles can evolve in A. fumigatus colonising the lungs, and strains with elevated minimum inhibitory concentrations (MIC) are often isolated from sputum samples of infected hosts undergoing azole antifungal therapy. Interestingly though, resistance strains of A. fumigatus have also been isolated from azole naive hosts, which has raised questions as to how and why azole resistance has evolved on these occasions. 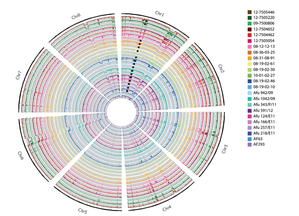 A. fumigatus whole genome alignment - Abdolrasouli et al 2015, mBio. A. fumigatus whole genome alignment - Abdolrasouli et al 2015, mBio. One leading theory suggests that resistance has evolved in the environment under the strong selection of azole fungicide use, which is often applied to agricultural and horticultural crops for the control of plant pathogenic fungi. There are currently two principal "environmental" mutations known to confer resistance to azole antifungals (TR34/L98H and TR46/Y121F/T289A). The two sets of mutations are found in cyp51A, the target of azole antifungals. They are comprised of a tandem repeat (TR) in the promoter region, that increases cyp51A expression, and at least one amino acid substitution that physically alters protein confirmation and decreases azole binding affinity. It is thought that normal dispersal of conidia via asexual reproduction has disseminated these resistance alleles into the wider population, with some resistance conidia inhaled into the lungs of immunocompromised hosts where a resistance phenotype is observed (despite no previous history of azole prescription). 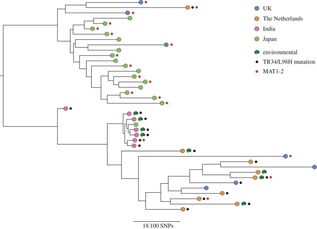 Phylogenetic analysis of A. fumigatus isolates - Abdolrasouli et al 2015, mBio. Phylogenetic analysis of A. fumigatus isolates - Abdolrasouli et al 2015, mBio. To test this theory, our lab applied the WGS approach; we used 24 genome sequences to infer the population structure of resistant and susceptible A. fumigatus isolates collected from both environmental and clinical settings worldwide. This preliminary study showed that sequencing resolution was high enough to identify isolates with and without the TR34/L98H mutation. It identified two distinct clades, which supports previous studies that described this level of structuring. And it showed that A. fumigatus is panmictic and recominogenic, and is probably able to adapt quickly to novel environmental conditions. Our aim now is to build on this previous study and include hundreds more genome sequences to better identify A. fumigatus population structure and evolutionary history in relation to the TR34/L98H and TR46/Y121F/T289A mutations. Hopefully this will enable us to provide a more concrete theory of how, why and when these mechanisms of azole resistance evolved. Work is well underway... please check back regularly for updates on our progress over the next few years! Human aspergillosis is an emerging infectious disease (EIDs) primarily caused by the ubiquitous, fungal saprophyte Aspergillus fumigatus. As an opportunistic human pathogen, A. fumigatus can causes a range of symptoms, from minor allergies to invasive aspergilloisis (IA), a life-threatening infection in immunocompromised hosts. In the environment, A. fumigatus plays a crucial role in the decomposition of organic material and can regularly be found in soil or on decaying plant material (e.g. compost or crop stubble).
Susceptible human hosts are infected with A. fumigatus through the inhalation of airborne condia, which are asexually produced spores dispersed on mass by the fungus. For established infections, azole antifungals, such as itraconazole, voriconazole and posconazole, are prescribed for management and prophylaxis of IA. However, the emergence of azole resistance in A. fumigatus has been identified globally and is now considered and evolving public health problem. |
Fisher Lab
|


